P3M
(Platform for
Manipulating
Molecular Interaction
Maps)
(Platform for Manipulating Molecular Interaction Maps)
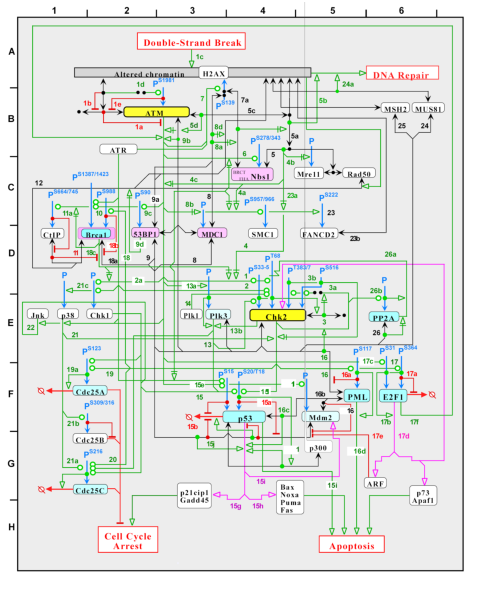
P3M is a prototypal system allowing one to reason about Molecular Interaction Maps (MIMs). The system takes as input files representing MIMs created by PathVisio, a free open-source biological pathway analysis software that allows one to draw biological pathways.
P3M is able to solve graph validation, graph querying and graph updating problems. Its mechanism relies upon the representation of the behaviour of the biological system by use of Linear Temporal Logic and its grounding into propositional logic. It was developped in a collaboration between the Institut de Recherche en Informatique de Toulouse (IRIT) and the Centre de Recherche en Cancérologie de Toulouse (CRCT).
The system is written in Objective Caml and interfaces with the C implementation of the Picosat solver library. The program compiles and runs under Linux and Cygwin, and should also work with MacOs.
A thorough description of P3M theoretical grounds and of the system itself can be found in the following draft:
Participants to the project are: Jean-Marc Alliot (IRIT), Marta Cialdea Mayer (Università degli Studi Roma Tre, Roma), Robert Demolombe (IRIT), Martín Diéguez (IRIT), Luis Fariñas del Cerro (IRIT), Gilles Favre (CRCT), Jean-Charles Faye (CRCT), Olivier Sordet (CRCT).
Download
P3M (Linux executable)
Sample input files
The sources of both the system and its documentation
Contacts
-
Jean-Marc Alliot.
Institut de recherche en informatique de Toulouse.
Toulouse University, Toulouse, France.
email: jean-marc.alliot@irit.fr -
Luis Fariñas Del Cerro,
Institut de recherche en informatique de Toulouse.
Toulouse University, Toulouse, France.
email: luis.farinas@irit.fr